
Anima software repository
We provide source code to our latest implementations of our research algorithms. It includes algorithms for registration, MS lesions segmentation, diffusion image processing, population statistics tools, quantitative MRI processing (relaxometry), image denoising. This software repository is intended to grow with new algorithms as they become available.
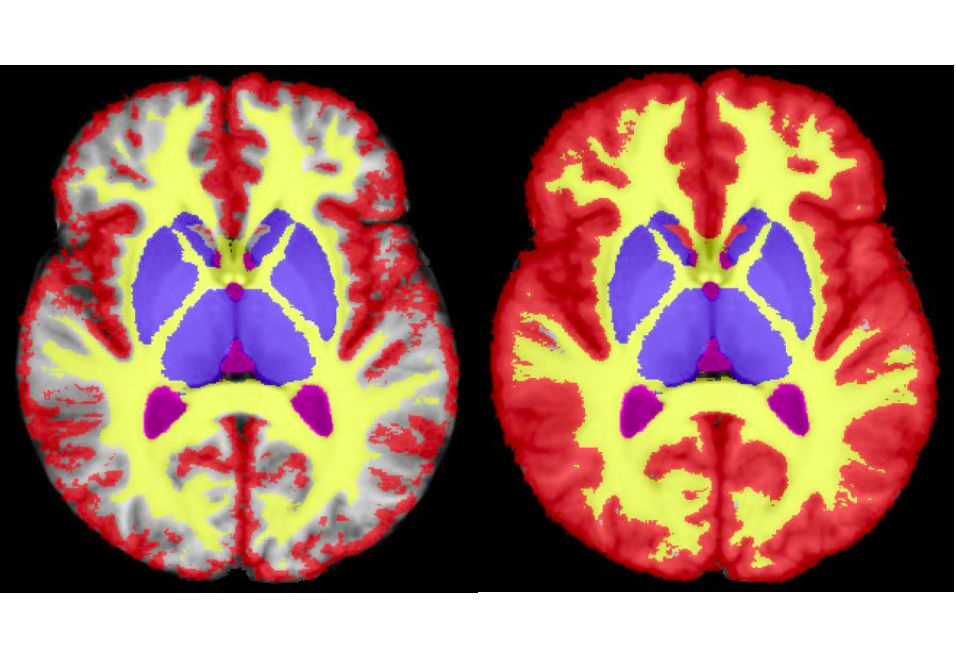
STAPLE variants implementation
We provide source code to our latest improvements on the original STAPLE algorithm. Namely, we distribute the MAP STAPLE algorithm as well as the continuous STAPLE algorithm on vector and scalar images. Soon, this will also include inferential uncertainty estimation as well as our recent local MAP STAPLE algorithm.
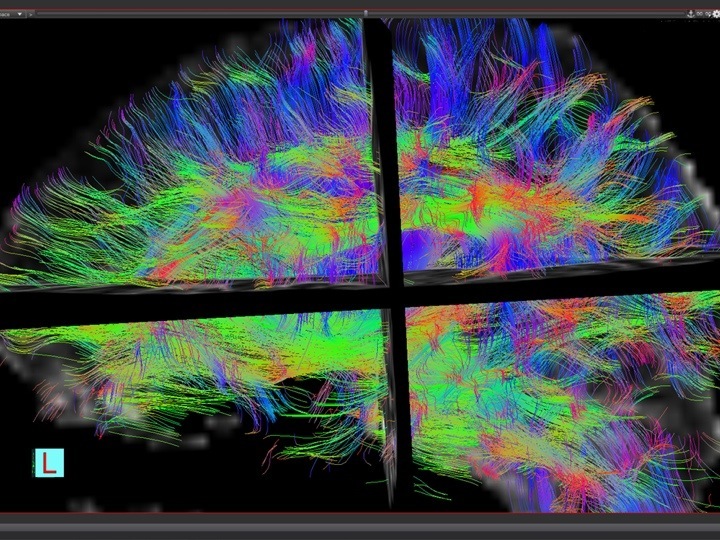
medInria
We are developing medInria, incorporating developments from major medical image processing teams at Inria. medInria is available open-source on Github from the medInria website. Some related packages from the Empenn team include the NL-means denoising algorithm, diffusion compartment imaging, and query/retrieve from a PACS or Shanoir servers.

LaTeX PhD and HDR template
This provides a template in French or English for a PhD thesis or HDR to be written in LaTeX. Some examples and scripts are included in the package.

Young and van Vliet Gaussian filter
As part of a collaborative work between Inria and Mauna Kea Technologies, the source code of a CPU and GPU implementation of the Young and van Vliet recursive Gaussian image filter has been made available. An Insight journal publication is available wit hthe source code, and the module has been integrated as a remote module inside ITK.